ABSTRACT Background: Bloodstream infections are a significant cause of community morbidity and mortality, and their microbiological patterns can vary regionally. In British Columbia, the Lower Mainland and Vancouver Island represent two major population centres with distinct demographic and exposure characteristics. Analysis of community blood culture data provides an opportunity to identify geographic differences in bloodstream pathogens and monitor regional antimicrobial resistance trends over time.
Polina Novoseltseva, BSc, MSc1 and Eugene Y. H. Yeung, MD, MSc, BSc (Pharm), FRCPC, FCCM, D(ABMM), DTM&H, CTH, CIC, LTC-CIP 1,2,3*
1 Faculty of Medicine, University of British Columbia, Vancouver, B.C., Canada
2 Faculty of Pharmaceutical Sciences, University of British Columbia, Vancouver, B.C., Canada
3 School of Medicine, Simon Fraser University, Surrey, B.C., Canada
*Corresponding author
Eugene Y.H. Yeung
LifeLabs, 7455 130 St.
Surrey, B.C., Canada V3W 1H8
Email: eugeney@doctors.org.uk
Article history:
Received 20 November 2025
Received in revised form 23 December 2025
Accepted 5 January 2026
ABSTRACT
Background: Bloodstream infections are a significant cause of community morbidity and mortality, and their microbiological patterns can vary regionally. In British Columbia, the Lower Mainland and Vancouver Island represent two major population centres with distinct demographic and exposure characteristics. Analysis of community blood culture data provides an opportunity to identify geographic differences in bloodstream pathogens and monitor regional antimicrobial resistance trends over time.
Methods: Blood culture data collected by LifeLabs British Columbia from 2020 to 2024 were analyzed, encompassing 744 community isolates: 620 from the Lower Mainland and 124 from Vancouver Island. Organisms were identified, and antimicrobial susceptibility results were compared between the two regions to assess regional variation in bloodstream infection patterns.
Results: Of all isolates, 62% were Gram-positive and 37% were Gram-negative; yeast accounted for 1% of isolates and were not analyzed further. In the Lower Mainland, Salmonella species (41% of gram-negative isolates) and Escherichia coli (E. coli) (32% of Gram-negative isolates) were the predominant Gram-negative organisms, whereas E. coli was most common on Vancouver Island (35% of Gram-negative isolates). Overall, Staphylococcus aureus (11%) and viridans group Streptococcus (22%) were the most frequently isolated Gram-positive organisms. Salmonella species and E. coli demonstrated reduced ciprofloxacin susceptibility (0% and 60%, respectively), while Salmonella remained universally susceptible to ceftriaxone (100%) and trimethoprim-sulfamethoxazole (100%).
Conclusion: This analysis provides region-specific surveillance data describing the distribution and antimicrobial susceptibility of community bloodstream pathogens in British Columbia. The identification of Salmonella predominance in the Lower Mainland and reduced ciprofloxacin susceptibility among E. coli highlights the importance of ongoing provincial surveillance. These findings contribute to a better understanding of regional epidemiology and inform infection control and antimicrobial resistance monitoring initiatives in community health settings.
KEYWORDS:
Antimicrobial susceptibility, community bacteremia, British Columbia
BACKGROUND
Bloodstream infections are a significant cause of morbidity and mortality worldwide; among hospitalized patients, a reported 90-day mortality rate of 36% has been observed (Santoro et al., 2020). In Canada, an Ontario-based study found that Escherichia coli (E. coli) bloodstream infection was associated with a 17.8% 90-day mortality among hospitalized patients between 2017 and 2020 (Daneman et al., 2023). In community-based care settings, timely identification of bloodstream pathogens remains a challenge because diagnostic results often become available only several days after specimen collection. To enhance regional understanding of community bacteremia, LifeLabs British Columbia (B.C.) developed a cumulative antimicrobial susceptibility testing (CAST) report encompassing all positive blood culture results from the Lower Mainland and Vancouver Island between 2020 and 2024.
A 2010-2014 LifeLabs study revealed that, in community blood cultures (n = 9,092), typhoidal Salmonella was as common as E. coli, a finding that differed significantly from hospital trends, which showed E. coli as the most common bloodstream
pathogen (Imperial et al., 2018; MacKinnon et al., 2021). This finding prompted a deeper analysis of the current regional bacteremia patterns and emerging resistance in community settings. Developed in accordance with Clinical and Laboratory Standards Institute (CLSI, 2025) guidance, our CAST provides longitudinal, region-specific data to inform public health surveillance and infection control programs. By tracking regional shifts in bloodstream pathogen distribution and susceptibility, this initiative enhances early detection of resistance trends and strengthens the evidence base for antimicrobial stewardship and regional planning initiatives in British Columbia.
Methods
Study design
This retrospective analysis used community blood culture data from LifeLabs British Columbia, Canada. The dataset included all positive blood cultures collected from non-hospitalized patients in the Lower Mainland and Vancouver Island between January 1, 2020, and December 31, 2024. Blood culture orders were primarily submitted by general practitioners in outpatient settings, and samples were processed at regional LifeLabs microbiology laboratories. Blood specimens were collected by trained phlebotomists using standard venipuncture techniques into aerobic and anaerobic culture bottles, which were transported under controlled conditions to regional laboratories. Bottles were incubated for up to five days in BACTEC automated culture systems, and those flagged as positive were subjected to Gram stain and Matrix-Assisted Laser Desorption Ionization–Time-of-Flight (MALDI-TOF) mass spectrometry for organism identification. Antimicrobial susceptibility testing (AST) was performed using the VITEK2 system, following Clinical and Laboratory Standards Institute (CLSI, 2025). When VITEK2 verification was unavailable, AST was completed using Kirby-Bauer disk diffusion or Etest methods. Duplicate isolates from the same patient and collection episode were excluded from AST analysis.
Data extraction and statistical analysis
All microbiological data were extracted from the LifeLabs Microbiology Electronic Worksheet System (Surrey, B.C., Canada). The dataset included organism identification and antimicrobial susceptibility profiles, with all patient identifiers removed prior to analysis. A total of 744 unique bloodstream isolates were included, with 620 from the Lower Mainland and 124 from Vancouver Island, representing bloodstream isolates from non-hospitalized patients processed at LifeLabs during the five-year period. Antimicrobial-susceptibility patterns were compared between the two regions using the chi-square test with GraphPad QuickCalcs software (Dotmatics). A p value < 0.05 was considered statistically significant.
Results
Microorganism distribution analysis
Among 744 pooled patient isolates from the Lower Mainland and Vancouver Island, 37% were Gram-negative organisms (n = 277), 62% were Gram-positive (n = 464), and 1% were yeast (n = 3).
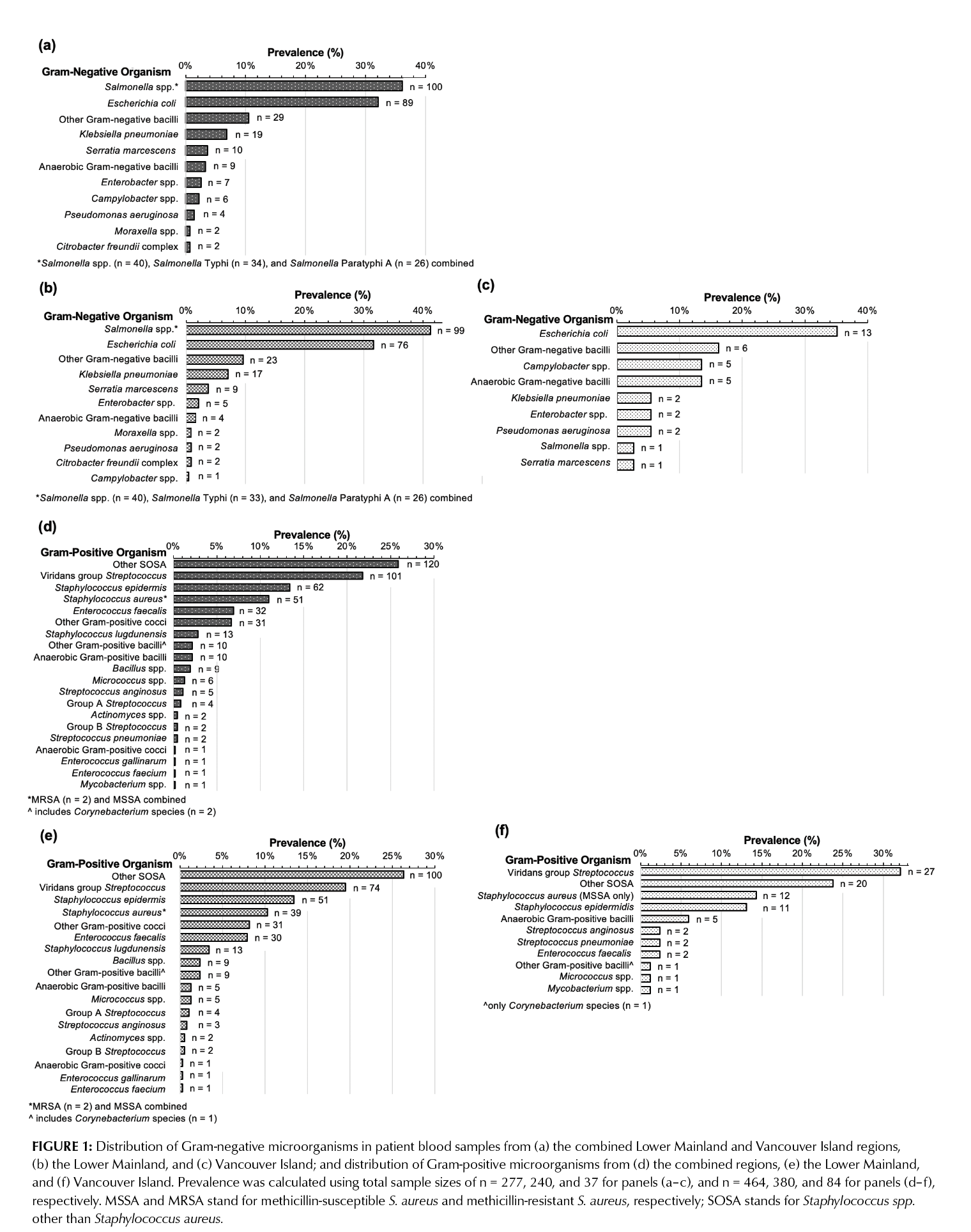
Within the Gram-negative group, Salmonella species accounted for 36% (n = 100) and E. coli for 32% (n = 89) of Gram-negative isolates. In the Lower Mainland, Salmonella species were the predominant Gram-negative pathogen, whereas on Vancouver Island, E. coli was most frequent (n = 13; 35% of Gram-negative isolates) (Figure 1a-c). Overall, Gram-negative organism distributions differed significantly between the two regions (p < 0.001).
The prevalence of Gram-positive microorganisms was similar between the Lower Mainland and Vancouver Island (Figure 1e and 1f); therefore, pooled results are presented (Figure 1d). S. aureus was the most frequent clinically significant isolate, representing 11% of Gram-positive cases (49 methicillin-sensitive S. aureus (MSSA) and two methicillin-resistant S. aureus (MRSA), both from the Lower Mainland). Other commonly isolated Gram-positive organisms included viridans group Streptococcus (22%), Staphylococcus epidermidis (13%), and other Staphylococcus species (26%).
Cumulative Antimicrobial Susceptibility Testing (CAST) trends
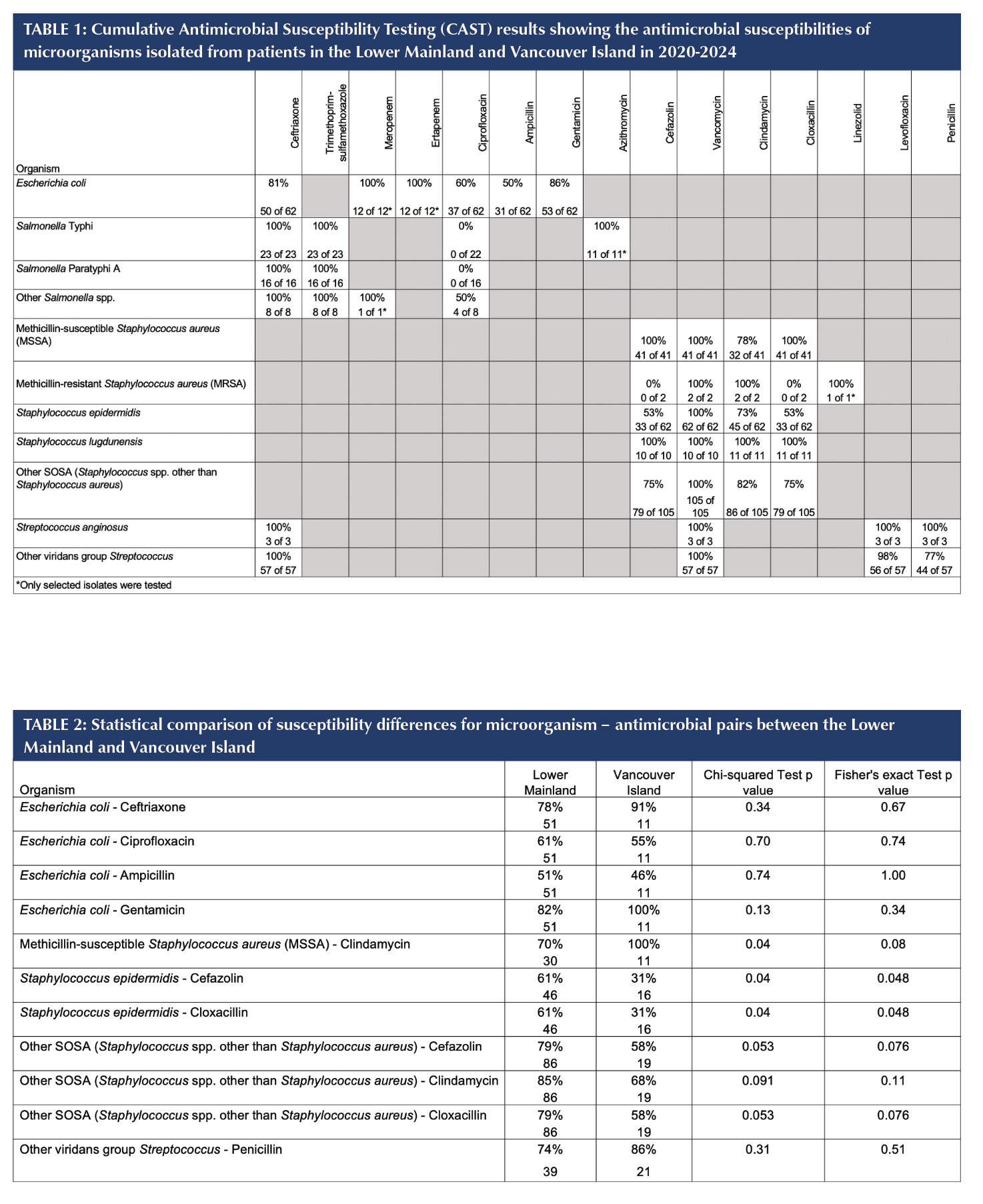
The pooled antimicrobial susceptibility profiles of all microorganisms identified in the Lower Mainland and Vancouver Island regions are summarized in Table 1.
Regional susceptibility differences were observed for select Gram-positive organisms (Table 2). MSSA isolates from the Lower Mainland demonstrated 70% susceptibility to clindamycin, compared with 100% on Vancouver Island (p = 0.04). For S. epidermidis, susceptibilities to cefazolin and cloxacillin were lower among isolates collected from residents of Vancouver Island (31%) than in the Lower Mainland (61%), and these differences were statistically significant (p = 0.04 and p = 0.048, respectively). Other coagulase-negative Staphylococcus (SOSA) isolates displayed a similar but non-significant trend (79% vs 58%).
Discussion
Gram-negative trends
Escherichia coli remained the predominant cause of community-acquired Gram-negative bacteremia in both Vancouver Island and the Lower Mainland regions, consistent with national data showing similar pre-COVID-19 distributions in Ontario populations (Hasan et al., 2024). Salmonella species surpassed E. coli in the Lower Mainland, a distinct regional difference that may reflect greater travel to Salmonella-endemic regions in Southeast Asia (Lo et al., 2025), a trend that may continue in the coming years.
Antimicrobial susceptibilities of Gram-negative organisms
Among Salmonella isolates from the Lower Mainland, all S. Typhi and S. Paratyphi A isolates were ciprofloxacin-resistant, while 50% of other Salmonella species showed resistance (Table 1). The single S. Typhi isolate from Victoria was likewise ciprofloxacin-resistant. Despite this, all Salmonella isolates were susceptible to ceftriaxone and trimethoprim-sulfamethoxazole.
For E. coli, pooled ceftriaxone susceptibility among tested isolates was 81%, with regional differences between the Lower Mainland (78%) and Vancouver Island (91%). These values align with broader North American trends showing increased ceftriaxone resistance since 2012, which is largely attributed by extended-spectrum beta-lactamase (ESBL) production in North America (Tamma et al., 2022). All E. coli isolates tested were susceptible to carbapenems.
Collectively, these data indicate notable antimicrobial resistance among community Gram-negative organisms, and highlight the importance of ongoing regional surveillance to detect early shifts in susceptibility.
Gram-positive trends
In both regions, Staphylococcus aureus accounted for approximately 11% of bloodstream isolates (Figure 1d), reinforcing its status as a leading cause of community-acquired bacteremia in Canada (Bai & Morris, 2019). Two of the
39 S. aureus cases from the Lower Mainland were methicillin-resistant (MRSA), while none were identified on Vancouver Island. Given the small number of MRSA cases in the Lower Mainland community, it is premature to draw conclusions about trends or sources of MRSA bacteremia.
Viridans group Streptococcus (22%) and coagulase-negative Staphylococcus (including S. epidermidis) represented 39% of Gram-positive isolates. When recovered from multiple culture bottles, viridans group Streptococcus and coagulase-negative Staphylococcus isolates are more likely to represent true bloodstream infections rather than contaminants (CLSI 2022). Viridans group Streptococcus remains an important cause of infective endocarditis in paediatric populations (Vicent et al., 2022) while S. epidermidis, part of the normal skin flora, is associated with device-related infections (Rogers et al., 2009). Three yeast bloodstream infections were identified exclusively in the Lower Mainland, consistent with their known occurrence
among immunocompromised or catheterized individuals (Pappas et al., 2018).
Overall, the distributions of S. aureus, viridans group Streptococcus, and SOSA were comparable between regions, suggesting similar Gram-positive bacteremia profiles in both communities and underscoring S. aureus as a persistent public health concern in B.C.
Antimicrobial susceptibilities of Gram-positive organisms
Methicillin-susceptible S. aureus (MSSA) isolates from the Lower Mainland showed clindamycin susceptibility of 70% in the Lower Mainland compared to 100% susceptibility among Vancouver Island isolates (Table 2). This finding parallels international reports indicating variable clindamycin activity against MSSA (Alvarez-Payares et al., 2022). All MSSA isolates were susceptible to vancomycin (Bai & Morris, 2019).
For S. epidermidis and other coagulase-negative Staphylococcus isolates, cefazolin and cloxacillin susceptibilities were 61% in the Lower Mainland and 31% on Vancouver Island, suggesting regional variation in beta-lactam susceptibility among Gram-positive isolates, emphasizing the value of region-specific surveillance (Table 2). These findings point to meaningful intraprovincial variation in antimicrobial susceptibility among Gram-positive isolates, emphasizing the value of region-specific surveillance data for interpreting local resistance trends.
Surveillance and public-health relevance
Although all isolates in this study originated from community settings, each represents a potentially invasive bloodstream infection with public-health importance. The observed regional differences in organism prevalence and antimicrobial resistance patterns underscore the value of maintaining timely communication between community laboratories and infection-control programs. Early regional data on Gram-stain results, available before full organism identification, are particularly useful for local epidemiological assessment, helping infection control teams and public health agencies identify emerging resistance clusters or outbreaks promptly.
Salmonella species accounted for 41% of Gram-negative bacteremia cases in the Lower Mainland compared to 3% on Vancouver Island, while E. coli remained the predominant Gram-negative isolate on Vancouver Island. These differences illustrate how geographic variation can influence early presumptive identification and inform local antimicrobial-stewardship decisions.
Contextualizing preliminary Gram-stain results within local resistance patterns can enhance surveillance of emerging antimicrobial resistance. Sharing these trends with community practitioners can improve awareness of locally prevalent pathogens and emerging susceptibility changes, supporting empiric treatment decisions. This approach can facilitate earlier recognition of shifts in community bacteremia trends, and support community-level antibiotic stewardship.
Study limitations
This study analyzed community-acquired bloodstream isolates collected between 2020 and 2024, and did not include hospital-derived isolates. Because of the limited sample size, data from the Lower Mainland and Vancouver Island were pooled for regional comparisons. In addition, susceptibility results for certain organism-antimicrobial combinations were based on fewer than 30 isolates and should be interpreted cautiously, in the context of published empirical therapy guidelines. Larger sample sizes would enable future studies to assess resistance trends over time. Certain antimicrobials, including azithromycin, were not assessed because CLSI interpretive breakpoints are not available for these organisms (Clinical and Laboratory Standards Institute, 2025). Despite these limitations, this study provides valuable regional insight into the epidemiology of community-acquired bloodstream infections and antimicrobial resistance trends in B.C.
Conclusion
This surveillance highlights distinct regional differences in bloodstream infection patterns in British Columbia. The predominance of Salmonella in the Lower Mainland and ongoing E. coli resistance to ceftriaxone and ciprofloxacin underscore the importance of maintaining community surveillance programs. Among Gram-positive organisms, S. aureus remains the most clinically significant pathogen, with MRSA restricted to the Lower Mainland during the study period. Continued regional monitoring is essential for tracking emerging resistance and supporting community practitioners in antimicrobial stewardship initiatives.
REFERENCES
Alvarez-Payares, J. C., Palacios M, J. E., De La Peña, E., Cabrera, H. B., Giraldo-Ramírez, S., Loaiza, M., Jaimes, F., Rodelo, J., & Ágamez-Gómez, J. (2022). Clindamycin Efficacy in Patients with Methicillin-Sensitive Staphylococcus aureus in a Fourth-Level Hospital in the City of Medellín. Cureus.
https://doi.org/10.7759/cureus.21124
Bai, A. D., & Morris, A. M. (2019). Management of Staphylococcus aureus bacteremia in adults. Canadian Medical Association Journal, 191(35), E967–E967.
https://doi.org/10.1503/cmaj.190363
Canadian Antimicrobial Resistance Surveillance System (CARSS). (2025, December 8). 2025 Highlights and Integrated Narrative: Antimicrobial Resistance (AMR) and Antimicrobial Use (AMU).
https://health-infobase.canada.ca/carss/#amr-
summary-findings
Canadian Nosocomial Infection Surveillance Program (CNISP). (2025, June 3). Healthcare-associated Infections and Antimicrobial Resistant Organisms. Government of Canada. https://health-infobase.canada.ca/cnisp/hai-aro-rates.html
Clinical and Laboratory Standard Institute (Ed.). (2022). CLSI M47: Principles and procedures for blood cultures (2. ed). Clinical and laboratory standard institute.
Clinical and Laboratory Standard Institute. (2025). CLSI M100: Performance standards for antimicrobial susceptibility testing (35th edition). Clinical and Laboratory Standards Institute.
Daneman, N., Fridman, D., Johnstone, J., Langford, B. J., Lee, S. M., MacFadden, D. M., Mponponsuo, K., Patel, S. N.,
Schwartz, K. L., & Brown, K. A. (2023). Antimicrobial resistance and mortality following E. coli bacteremia. eClinicalMedicine, 56, 101781.
https://doi.org/10.1016/j.eclinm.2022.101781
Dotmatics. (n.d.). Analyze a 2x2 contingency table. Retrieved May 24, 2025, from
https://www.graphpad.com/quickcalcs/contingency1/
Hasan, M. R., Vincent, Y. M., Leto, D., & Almohri, H. (2024). Microbiology of bloodstream infections in Ontario, Canada during COVID-19 pandemic. Canada Communicable Disease Report, 50(3/4), 114–118.
https://doi.org/10.14745/ccdr.v50i34a05
Imperial, M. R., Woo, L., Whellams, D., Liao, R. S., Reyes, R.,
& Kelly, M. (2018). A Retrospective Analysis from 2010–2014 Shows that Typhoidal Salmonella is as
Likely as Eschericia coli to be Recovered from Blood
Culture in a Large Outpatient Laboratory in the South
Fraser Region of British Columbia. Journal of the
Association of Medical Microbiology and Infectious
Disease Canada., 3(S1), 1–127.
https://doi.org/10.3138/jammi.3.suppl.01
Lo, C. K.-F., Mok, M., Schonhofer, C., Afra, K., & Masud, S. (2025). Current Antimicrobial Susceptibility Trends and Clinical Outcomes of Typhoidal Salmonella in a Large Health Authority in British Columbia, Canada. Tropical Medicine and Infectious Disease, 10(4), 108.
https://doi.org/10.3390/tropicalmed10040108
MacKinnon, M. C., McEwen, S. A., Pearl, D. L., Parfitt, E. C., Pasquill, K., Steele, L., & Laupland, K. B. (2021). Escherichia coli bloodstream infections in the western interior of British Columbia, Canada: A population-based cohort study. Epidemiology and Infection, 149, e195.
https://doi.org/10.1017/S0950268821001874
Pappas, P. G., Lionakis, M. S., Arendrup, M. C., Ostrosky-Zeichner, L., & Kullberg, B. J. (2018). Invasive candidiasis. Nature Reviews Disease Primers, 4(1), 18026.
https://doi.org/10.1038/nrdp.2018.26
Rogers, K. L., Fey, P. D., & Rupp, M. E. (2009). Coagulase-Negative Staphylococcal Infections. Infectious Disease Clinics of North America, 23(1), 73–98.
https://doi.org/10.1016/j.idc.2008.10.001
Santoro, A., Franceschini, E., Meschiari, M., Menozzi, M., Zona, S., Venturelli, C., Digaetano, M., Rogati, C., Guaraldi, G., Paul, M., Gyssens, I. C., & Mussini, C. (2020). Epidemiology and Risk Factors Associated with Mortality in Consecutive Patients with Bacterial Bloodstream Infection: Impact of MDR and XDR Bacteria. Open Forum Infectious Diseases, 7(11), ofaa461.
https://doi.org/10.1093/ofid/ofaa461
Tamma, P. D., Komarow, L., Ge, L., Garcia-Diaz, J., Herc, E. S., Doi, Y., Arias, C. A., Albin, O., Saade, E., Miller, L. G., Jacob, J. T., Satlin, M. J., Krsak, M., Huskins, W. C., Dhar, S., Shelburne, S. A., Hill, C., Baum, K. R., Bhojani, M., … for the Antibacterial Resistance Leadership Group. (2022). Clinical Impact of Ceftriaxone Resistance in Escherichia coli Bloodstream Infections: A Multicenter Prospective Cohort Study. Open Forum Infectious Diseases, 9(11), ofac572.
https://doi.org/10.1093/ofid/ofac572
Tchesnokova, V., Larson, L., Basova, I., Sledneva, Y., Choudhury, D., Solyanik, T., Heng, J., Bonilla, T. C., Pham, S., Schartz, E. M., Madziwa, L. T., Holden, E., Weissman, S. J., Ralston, J. D., & Sokurenko, E. V. (2023). Increase in the community circulation of ciprofloxacin-resistant Escherichia coli despite reduction in antibiotic prescriptions. Communications Medicine, 3(1), 110.
https://doi.org/10.1038/s43856-023-00337-2


